Research Programs

Interdisciplinary Research Units
Explore our unique network of Organized Research Units and Special Research Programs that bring together experts from different field of study to address some of society’s most complex issues.

Core Facilities and Services
Discover how our Core Facilities provide access to shared resources, such as technologies, equipment, services, and expert consultation to ensure a world-call research infrastructure.

UC Davis Grand Challenges
Learn about how UC Davis is catalyzing our campus community to go beyond team science to holistically tackle the Earth’s most daunting challenges.
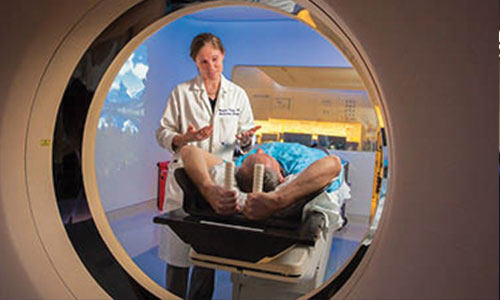
Clinical Research
UC Davis conducts more than 1,000 clinical research studies annually, including basic science, translational and clinical trial research — all with the goal of bringing new, effective and safe treatments to patients.

Colleges and Schools Research
Explore the research being conducted by faculty within our four colleges and six professional schools that drives advancements in each specific field of study.

Faculty Expertise
Find experts in specific areas of study using one of our publicly available search portals.

